Single-subject Proteomics Analysis Workflows
Single-subject Proteomic Signatures in Alzheimer’s Disease Reflect Clinical Phenotypes
and Distinguish Asymptomatic from Symptomatic Cases
Authors:
Avijit Podder¹, Yi Juin Liew¹, Greg A. Cary², Gregory W. Carter¹², Asli Uyar¹
Affiliations:
¹ The Jackson Laboratory for Genomic Medicine, Farmington, CT, USA
² The Jackson Laboratory, Bar Harbor, ME, USA
Corresponding authors:
Gregory W. Carter (gregory.carter@jax.org); Asli Uyar (asli.uyar@jax.org)
This site provides the rendered HTML workflows and supporting code for the analyses
reported in the manuscript. The reports cover TMT proteomics processing,
single-subject differential expression analysis, pathway- and subdomain-level
profiling, graph-based clustering, and clinical correlation analyses.
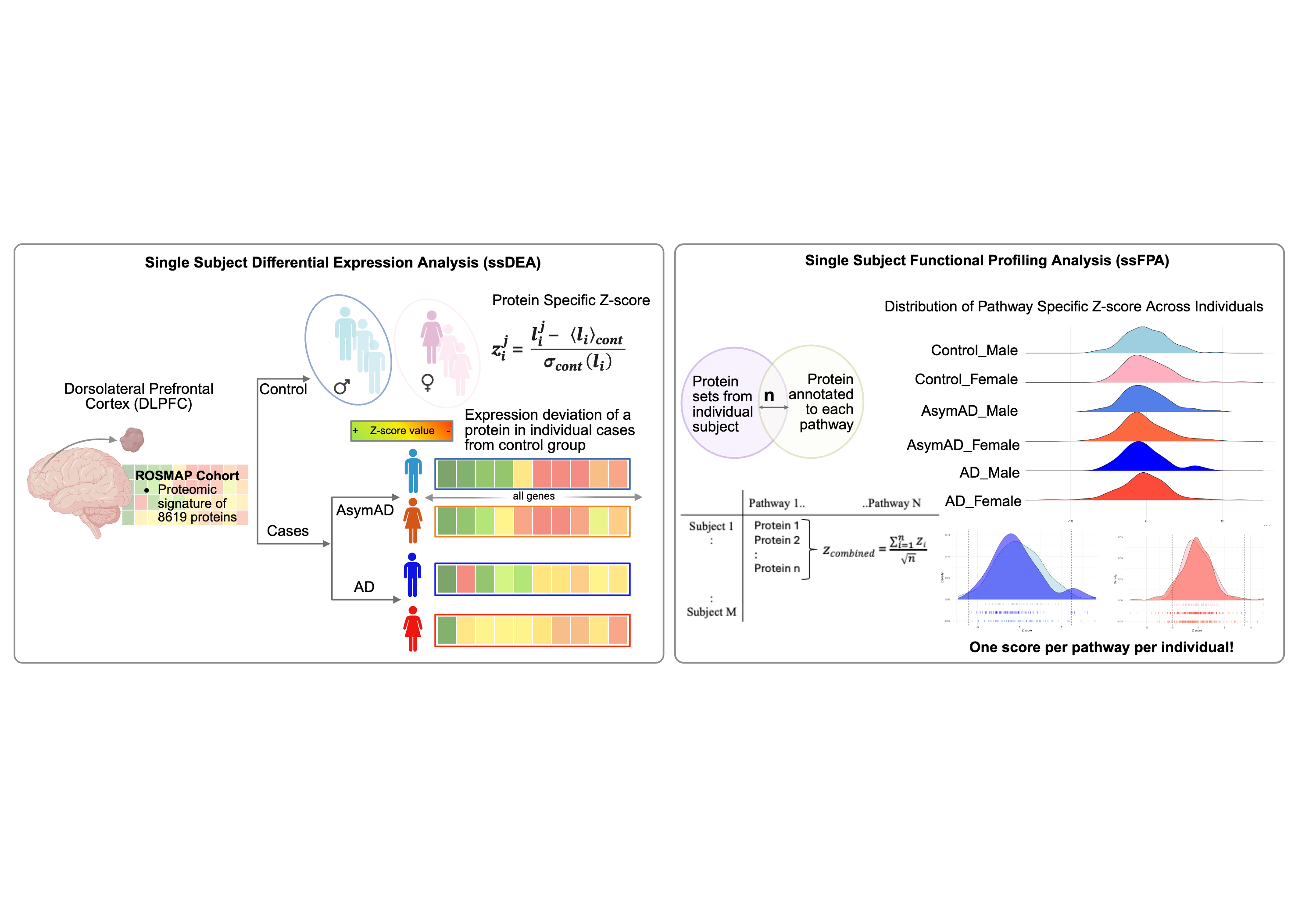
Schematic overview of the proteomics analysis pipeline used in this study.
Analysis reports

